Filipe M. Sousa, Luís M. P. Lima, Clément Arnarez, Manuela M. Pereira, and Manuel N. Melo
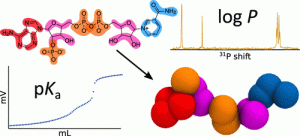
Nucleotides are structural units relevant not only in nucleic acids but also as substrates or cofactors in key biochemical reactions. The size- and timescales of such nucleotide–protein interactions fall well within the scope of coarse-grained molecular dynamics, which holds promise of important mechanistic insight. However, the lack of specific parameters has prevented accurate coarse-grained simulations of protein interactions with most nucleotide compounds. In this work, we comprehensively develop coarse-grained parameters for key metabolites/cofactors (FAD, FMN, riboflavin, NAD, NADP, ATP, ADP, AMP, and thiamine pyrophosphate) in different oxidation and protonation states as well as for smaller molecules derived from them (among others, nicotinamide, adenosine, adenine, ribose, thiamine, and lumiflavin), summing up a total of 79 different molecules. In line with the Martini parameterization methodology, parameters were tuned to reproduce octanol–water partition coefficients. Given the lack of existing data, we set out to experimentally determine these partition coefficients, developing two methodological approaches, based on 31P-NMR and fluorescence spectroscopy, specifically tailored to the strong hydrophilicity of most of the parameterized compounds. To distinguish the partition of each relevant protonation species, we further potentiometrically characterized the protonation constants of key molecules. This work successfully builds a comprehensive and relevant set of computational models that will boost the biochemical application of coarse-grained simulations. It does so based on the measurement of partition and acid–base physicochemical data that, in turn, covers important gaps in nucleotide characterization.
Doi: 10.1021/acs.jcim.0c01077
Cited as: Sousa FM, Lima LMP, Arnarez C, Pereira MM, Melo MN (2021) Coarse-Grained Parameterization of Nucleotide Cofactors and Metabolites: Protonation Constants, Partition Coefficients, and Model Topologies. J. Chem. Inf. Model. 61(1), 335–346; https://doi.org/10.1021/acs.jcim.0c01077.