Beatriz Ramos, Stephen V. Gordon, and Mónica V. Cunha
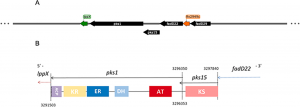
One of the most important and exclusive characteristics of mycobacteria is their cell wall. Amongst its constituent components are two related families of glycosylated lipids, diphthioceranates and phthiocerol dimycocerosate (PDIM) and its variant phenolic glycolipids (PGL). PGL have been associated with cell wall impermeability, phagocytosis, defence against nitrosative and oxidative stress and, intriguingly, biofilm formation. In bacteria from the Mycobacterium tuberculosis complex (MTBC), the biosynthetic pathway of the phenolphthiocerol moiety of PGL depends upon the expression of several genes encoding type I polyketide synthases (PKS), namely ppsA-E and pks15/1 which constitute the PDIM + PGL locus, and that are highly conserved in PDIM/PGL-producing strains. Consensus has not been achieved regarding the genetic organization of pks15/1 locus and knowledge is lacking on its transcriptional signature. Here we explore publicly available datasets of transcriptome data (RNA-seq) from more than 100 MTBC experiments in 40 growth conditions to outline the transcriptional structure and signature of pks15/1, using a differential expression approach to infer the regulatory patterns involving these and related genes. We show that pks1 expression is highly correlated with fadD22, Rv2949c, lppX, fadD29 and, also, pks6 and pks12, with the first three putatively integrating into a polycistronic structure. We evidence dynamic transcriptional heterogeneity within the genes involved in phenolphtiocerol and phenolic glycolipid production, most exhibiting up-regulation upon acidic pH and antibiotic exposure and down-regulation under hypoxia, dormancy, and low/high iron concentration. We finally propose a model based on transcriptome data in which σD positively regulates pks1, pks15 and fadD22, while σB and σE factors exert negative regulation at an upper level.
Doi: 10.1371/journal.pone.0229700
Cited as: Ramos B, Gordon SV, Cunha MV (2020) Revisiting the expression signature of pks15/1 unveils regulatory patterns controlling phenolphtiocerol and phenolglycolipid production in pathogenic mycobacteria. PLOS ONE 15(5): e0229700; https://doi.org/10.1371/journal.pone.0229700.